Products
Based on our experience, the kits for DNA extraction are recommended as follows.
|
Item |
Recommendations |
|
|
Extraction kit |
gDNA |
DNeasy Blood & Tissue Kit (Qiagen, Cat # 69504 & 69506)
TIANamp Genomic DNA Kit (TIANGEN, Cat # DP304) |
|
cfDNA |
QIAamp Circulating Nucleic Acid Kit (Qiagen, Cat # 55114) |
|
|
FFPE gDNA |
QIAamp DNA FFPE Tissue Kit (Qiagen, Cat # 56404) |
|
|
Dissolvent |
Nuclease-free water, 0.1 x TE (low TE), 10 mM Tris-HCl (pH 8.0-8.5) or Buffer provided in extraction kit should be used to dissolve DNA. |
|
This kit is compatible with types of dsDNA inputs, including DNA extracted from blood, tissues, FFPE and body fluid. This kit can be used to construct libraries for dsDNA input ranging from 1 to 500 ng.
For samples with suboptimal quality such as FFPE tissue samples, careful quality control (QC) and stratification on DNA integrity is highly recommended.
For common usage, the QC metrics are recommended as follows.
Item
Recommendations
Quantification
NanoDrop (based on the spectrophotometer principle), Qubit® or PicoGreen® (based on double-stranded DNA fluorescent dye).
Quality
OD260/OD280 = 1.8 - 2.0; OD260/OD230 = 2.0 - 2.5.
OD260/OD280 > 1.9 indicates RNA contamination while < 1.6 indicates protein, phenol or other contamination;
OD260/OD230 < 2.0 indicates carbohydrates, salts or organic solvent contamination.
Fragment distribution
gDNA
A clear band > 15 kb when running on a 1% agarose gel.
cfDNA
A peak at ~ 160 bp followed by a peak at ~ 320 bp on Bioanalyzer or Bioptic (Qsep).
FFPE gDNA
DNA integrity assessment by running on a 1% agarose gel or Bioanalyzer.
No. PCR with indexing primers is necessary to generate full-length library.
The kit is compatible with ultrasonic and enzymatic fragmentation methods. For enzymatic fragmentation, NEBNext® dsDNA Fragmentase® (NEB; Cat M0348) or Custom 5X DNA Shearing Mix (Qiagen; Cat CM0162) is recommended; For ultrasonic fragmentation, it is recommended to use Covaris (M220 Focused-ultrasonicator, 50 μL interruption tube) or similar.
For different applications, adapter module is recommended as follows.
|
Application |
Recommended adapter module |
Optional |
|
Whole-genome, metagenomic and targeted capture sequencing |
Universal Adapter (SI) Module |
BMI Adapter (SI) Module |
|
RNA sequencing |
BMI Adapter (SI) Module |
Universal Adapter (SI) Module |
|
Low frequency mutation detection (e.g. ≤ 1% variant frequency) |
BMI Adapter (SI) Module |
- |
The following tips help to improve the cleanup efficiency.
1) never overdry the beads;
2) increase incubation time;
3) increase the volume of magnetic beads (NOT suitable for size selection step);
4) use freshly prepared 80% ethanol.
Adapter: insert molar ratio is essential. Excessive adapter will result in residues of adapter or adapter dimers. Insufficient adapter will affect ligation efficiency and therefore reduce library yield. For universal adapters, please refer to the listed table to calculate the dilution fold of adapters so as to achieve the best yield.
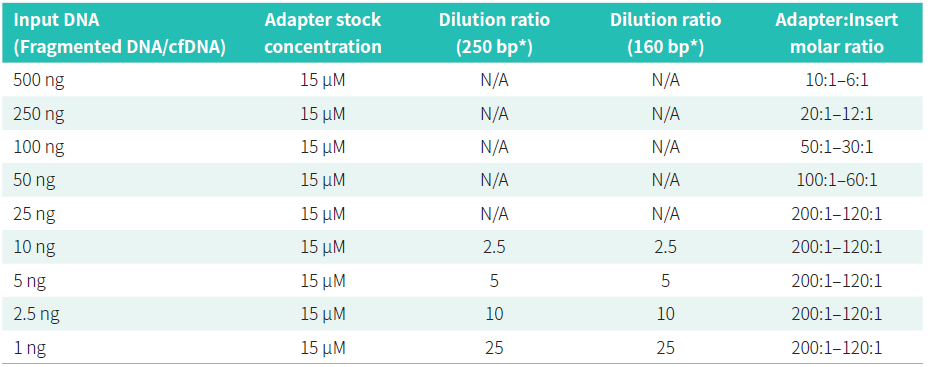
* ~ 250 bp is commonly used for gDNA fragmentation, and ~ 160 bp is peak size observed for cfDNA in plasma. Other recommendations for handling adapters are as follows.
• Diluent: Use TE Solution in the kit to dilute adapters. Diluted adapters are not recommended for long-term storage;
• Storage: Aliquot to avoid repeated freezing and thawing.
Ligation Buffer, DNA Ligase and End Repair & A-Tailing Enzyme are relatively viscous. It is recommended to use low-absorption tips and pipetting slowly. If residue remains in tip, repeatedly pipette or use the reverse pipetting.
We provided a total solution for targeted capture sequencing. The library kit is compatible with a variety of targeted capture systems, such as SureSelect (Agilent) and NimbleGen™ SeqCap™ EZ (Roche).