Products
If the nucleic acid samples contain high concentrations of EDTA or have an excessively high pH, it can affect the efficiency of DNA digestion. It is recommended to purify nucleic acid samples using 2X NadPrep SP Beads, replacing the elution reagent with Nuclease Free Water before proceeding with library preparation.
√ If the sample is a co-extracted total nucleic acid, quantification of RNA and DNA should be performed separately using Qubit, and library amplification should be carried out according to the recommended cycles in the manual.
√ During the experiment, RNase-free consumables must be used, and the operating environment should be free of RNase contamination, as this can affect the efficiency of RNA reverse transcription.
A: This solution is suitable for RNA samples from a wide range of cell and tissue sources with varying degradation levels. However, it is not recommended for severely degraded FFPE samples (DV 200 < 20%).
A: Yes, it is. This solution can be integrated with RNA library preparation kits for the preparation of total RNA sequencing (RNA-Seq) libraries. If you choose a third-party RNA library preparation kit, we recommend using NEBNext® Ultra™ II RNA Library Prep Kit for Illumina® (NEB, Cat # E7770L) or the KAPA RNA HyperPrep Kit (Roche, Cat # KK8541). For the use of RNA library preparation kits from other brands, please consult technical support from Nanodigmbio with support@njnad.com.
A: This solution contains a large amount of rRNA ssDNA blocking reagent. When used in conjunction with the NadPrep Total RNA-To-DNA Module, there may be a slight residual of ssDNA in the purified dsDNA product. When quantifying dsDNA using a nucleic acid fluorescence quantifier, this residual ssDNA can influence the results, leading to a slightly higher yield. However, the remaining rRNA ssDNA blocking reagent does not affect subsequent experiments. We recommend appropriate adapter dilution factors and PCR amplification cycles for different different types of RNA and different input amounts.
The kit can be used after being placed on the table at room temperature overnight without UV light irradiation. NadPrep EZ DNA Library Preparation Kit v2 hasa stable performance even at room temperature 12 h.
NadPrep EZ DNA Library Preparation Kit v2 is recommended to be frozen and thawed within 15 times. In order to ensure the data quality, it is recommended to use it in sub packaging. Please mix well before sub packaging.
In order to ensure the data quality, it is best to prepare the library within the scope recommended in the user manual. If there are special experimental needs, it can also be adjusted appropriately. The libraries prepared fromdifferent input DNA areshown as follows:
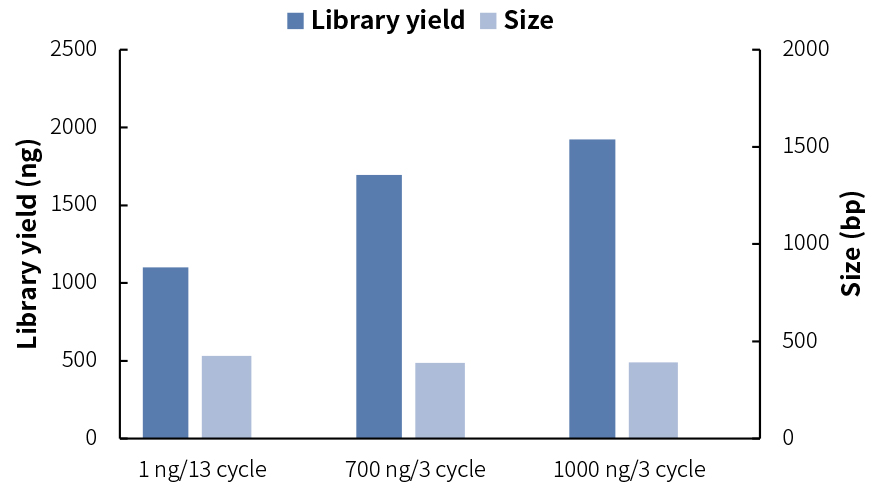
Experimental conditions: Human Genomic DNA Male (Promega, catalog # G1471) was fragmented with different input DNA, with 25 min of the enzymatic digestion time. NadPrep Universal Stubby Adapter (UDI) Module was used together for library preparation with single-sided selection mode. The library yield and size meet the requirements of sequencing.
|
Size Selection |
Applicable Situation |
Advantage |
Disadvantage |
|
Step 1
Optional steps:
Double-sided size selection after fragmentation |
Sufficient DNA input;
Broad size distribution of DNA
fragments after fragmentation |
Narrow size distribution,
high ligation efficiency and
uniform sequencing data |
Loss of sample |
|
Step 4
Optional steps:
Double-sided size selection after ligation |
Sufficient DNA input;
Broad size distribution of
libraries after adapter ligation |
Narrow size distribution,
Uniform sequencing data |
Strict double-sized
selection parameters
based on library size |
- Loss caused by double-sided selection after fragmentation
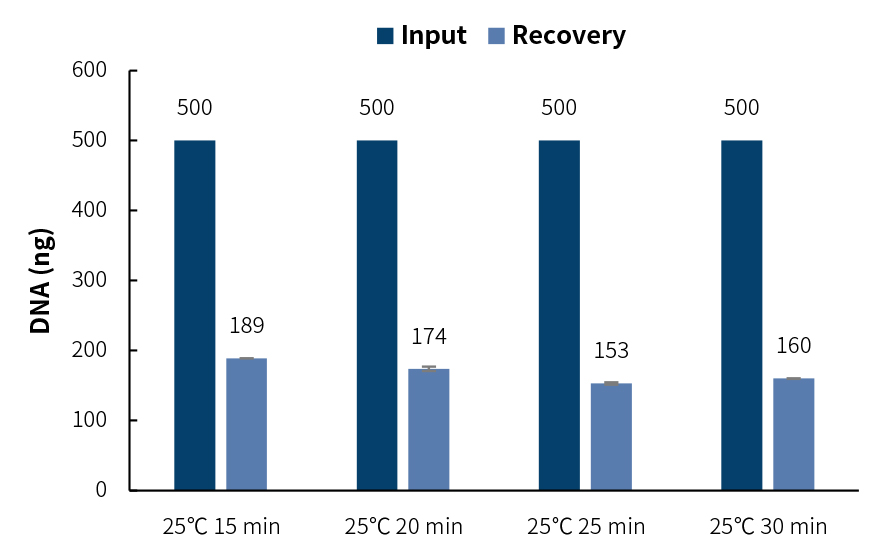
Experimental conditions: 500 ng of Human Genomic DNA Male (Promega, catalog # G1471) was fragmented by using different enzymatic digestion times, and the procedures were conduct at 25℃ for 15, 20, 25 and 30min, respectively, then65℃ for 30 min. At the end of the fragmentation procedures, according to Appendix II 1.1 of the user manual, double-sided selection was used. After selection, Nuclease Free water was used to elute for subsequent adapter ligation.
This step of selection can make the fragmented samples more concentrated, thus leading to high efficiency of subsequent adapter ligation, but the sample loss is relatively high (about 60-70%), which is suitable for the library preparation of sufficient DNA. It is recommended that input DNA is ≥200 ng
- Loss caused by double-sided selection after ligation
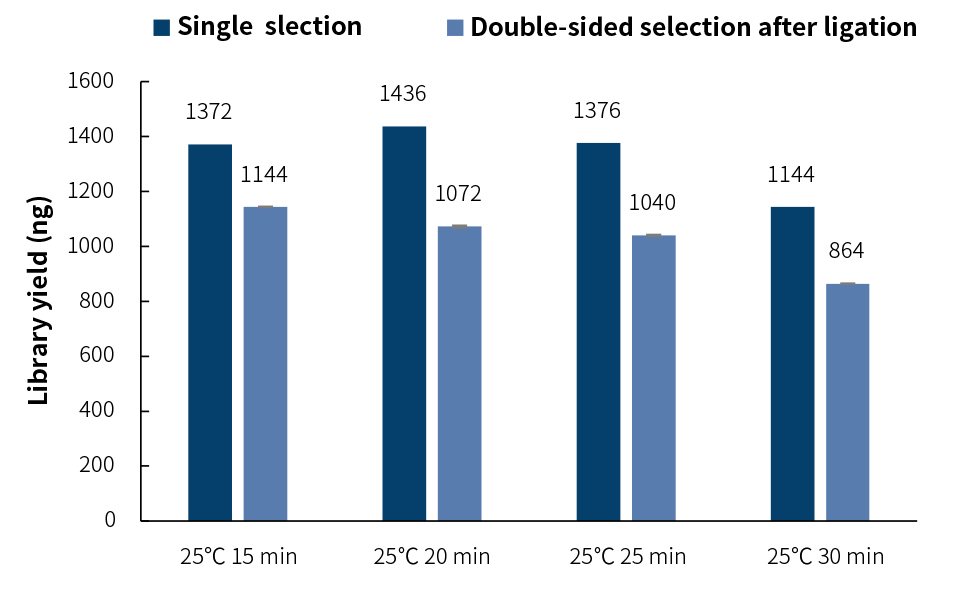
Experimental conditions: 100 ng of Human Genomic DNA Male (Promega, catalog # G1471) was fragmented by using different enzymatic digestion times. After the adapter ligation was
completed, the single screen mode and double screen mode were used respectively (see Appendix II 1.2 of the user manual for details).
The library yield after double-sided selection is 15-25% loss compared with single selection. But it is not recommended for low-quality DNA or input DNA is ≤50 ng.
|
Brand |
Kit Name |
Catalog |
Elution Solvent Name |
Elution Solvent Composition |
|
Promega |
MagneSil Blood Genomic, Max Yield System |
MD1360 |
Elution Buffer |
(10 mM Tris-Cl, 1 mM EDTA, pH 8.0) |
|
QIAGEN |
QIAamp DNA FFPE |
56404 |
Buffer ATE |
(10 mM Tris-Cl, 1 mM EDTA, pH 8.0) |
|
GeneRead DNA FFPE Kit |
180134 |
Buffer ATE |
(10 mM Tris-Cl, 1 mM EDTA, pH 8.0) |
|
|
DNeasy Blood & Tissue Kit |
69506 |
Buffer EB |
10 mM Tris-Cl, pH 8.5 |
|
|
Thermo Fisher Scientific |
PureLink Genomic DNA Mini Kit |
K182001 |
Elution Buffer |
10 mM Tris-HCl, pH 9.0, 0.1 mM EDTA |
|
Tiangen Biotech (Beijing) Co., Ltd. |
Magnetic beads-based genomic DNAextraction kit |
DP705 |
Buffer TB |
10 mM Tris-Cl, pH 8.0 |
|
Blood/cell/tissue genomic DNA extraction kit |
DP304 |
Buffer TE |
(10 mM Tris-Cl, 1 mM EDTA, pH 8.0) |
|
|
Paraffin embedded tissue DNA extraction kit (centrifugal column type) |
DP331 |
Buffer TE |
(10 mM Tris-Cl, 1 mM EDTA, pH 8.0) |
|
|
Cwbio |
Blood Genomic DNA Midi Kit (1-5 mL) |
CW0541 |
Buffer GE |
10 mM Tris,0.1 mM EDTA(pH 8.5) |
|
FlexGen Blood DNA Kit (0.1-20 mL) |
CW0544 |
Buffer GE |
10 mM Tris,0.1 mM EDTA(pH 8.5) |
|
|
FFPE DNA Kit |
CW0547 |
Buffer GE |
10 mM Tris,0.1 mM EDTA(pH 8.5) |